Posted by Jinsung Yoon and Sercan O. Arik, Research Scientists, Google Research, Cloud AI Team 
Analysis of Electronic Health Records (EHR) has a tremendous potential for enhancing patient care, quantitatively measuring performance of clinical practices, and facilitating clinical research. Statistical estimation and machine learning (ML) models trained on EHR data can be used to predict the probability of various diseases (such as diabetes), track patient wellness, and predict how patients respond to specific drugs. For such models, researchers and practitioners need access to EHR data. However, it can be challenging to leverage EHR data while ensuring data privacy and conforming to patient confidentiality regulations (such as HIPAA).
Conventional methods to anonymize data (e.g., de-identification) are often tedious and costly. Moreover, they can distort important features from the original dataset, decreasing the utility of the data significantly; they can also be susceptible to privacy attacks. Alternatively, an approach based on generating synthetic data can maintain both important dataset features and privacy.
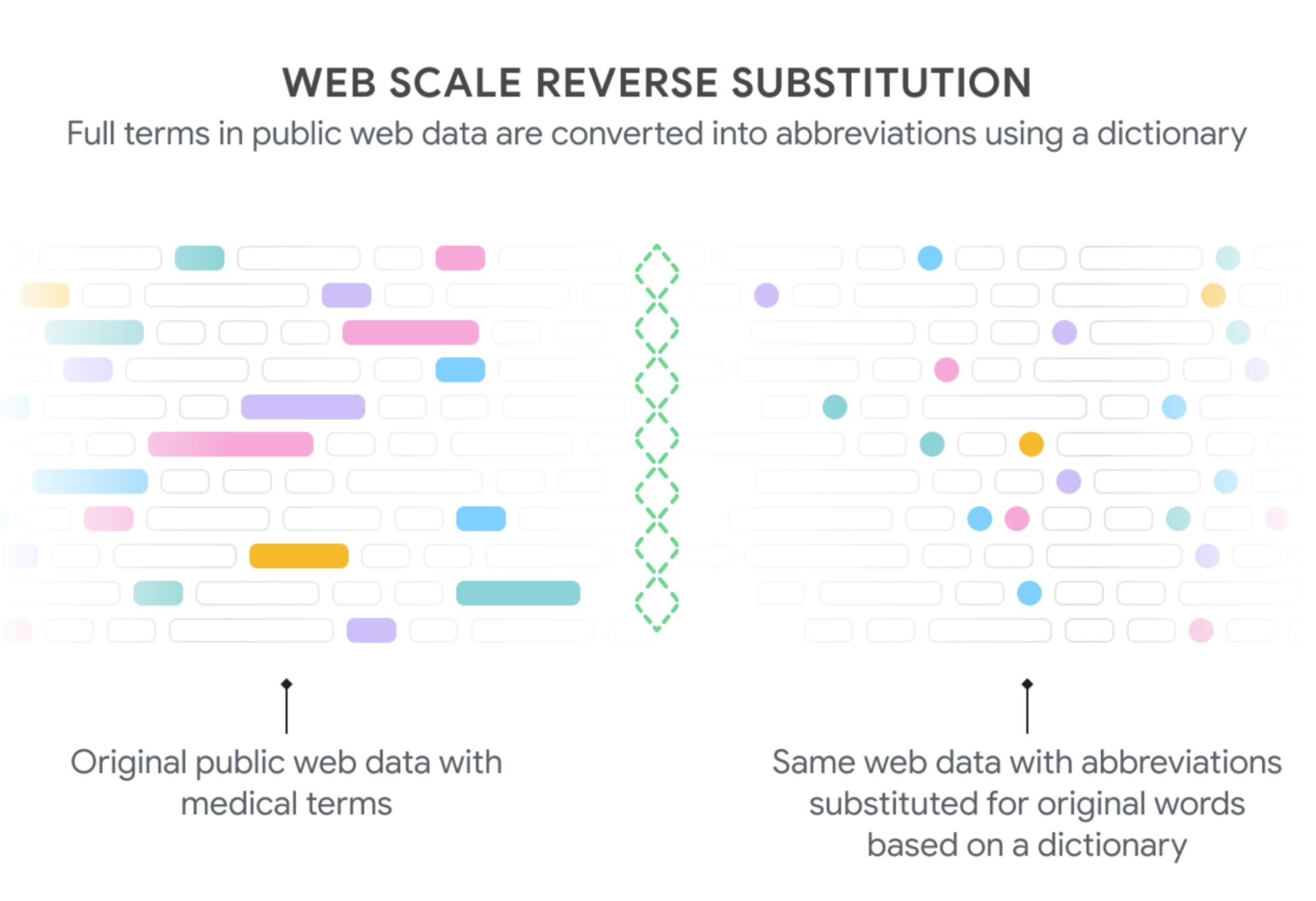
To that end, we propose a novel generative modeling framework in “EHR-Safe: Generating High-Fidelity and Privacy-Preserving Synthetic Electronic Health Records". With the innovative methodology in EHR-Safe, we show that synthetic data can satisfy two key properties: (i) high fidelity (i.e., they are useful for the task of interest, such as having similar downstream performance when a diagnostic model is trained on them), (ii) meet certain privacy measures (i.e., they do not reveal any real patient's identity). Our state-of-the-art results stem from novel approaches for encoding/decoding features, normalizing complex distributions, conditioning adversarial training, and representing missing data.
 |
| Generating synthetic data from the original data with EHR-Safe. |
Challenges of Generating Realistic Synthetic EHR Data
There are multiple fundamental challenges to generating synthetic EHR data. EHR data contain heterogeneous features with different characteristics and distributions. There can be numerical features (e.g., blood pressure) and categorical features with many or two categories (e.g., medical codes, mortality outcome). Some of these may be static (i.e., not varying during the modeling window), while others are time-varying, such as regular or sporadic lab measurements. Distributions might come from different families — categorical distributions can be highly non-uniform (e.g., for under-represented groups) and numerical distributions can be highly skewed (e.g., a small proportion of values being very large while the vast majority are small). Depending on a patient's condition, the number of visits can also vary drastically — some patients visit a clinic only once whereas some visit hundreds of times, leading to a variance in sequence lengths that is typically much higher compared to other time-series data. There can be a high ratio of missing features across different patients and time steps, as not all lab measurements or other input data are collected.
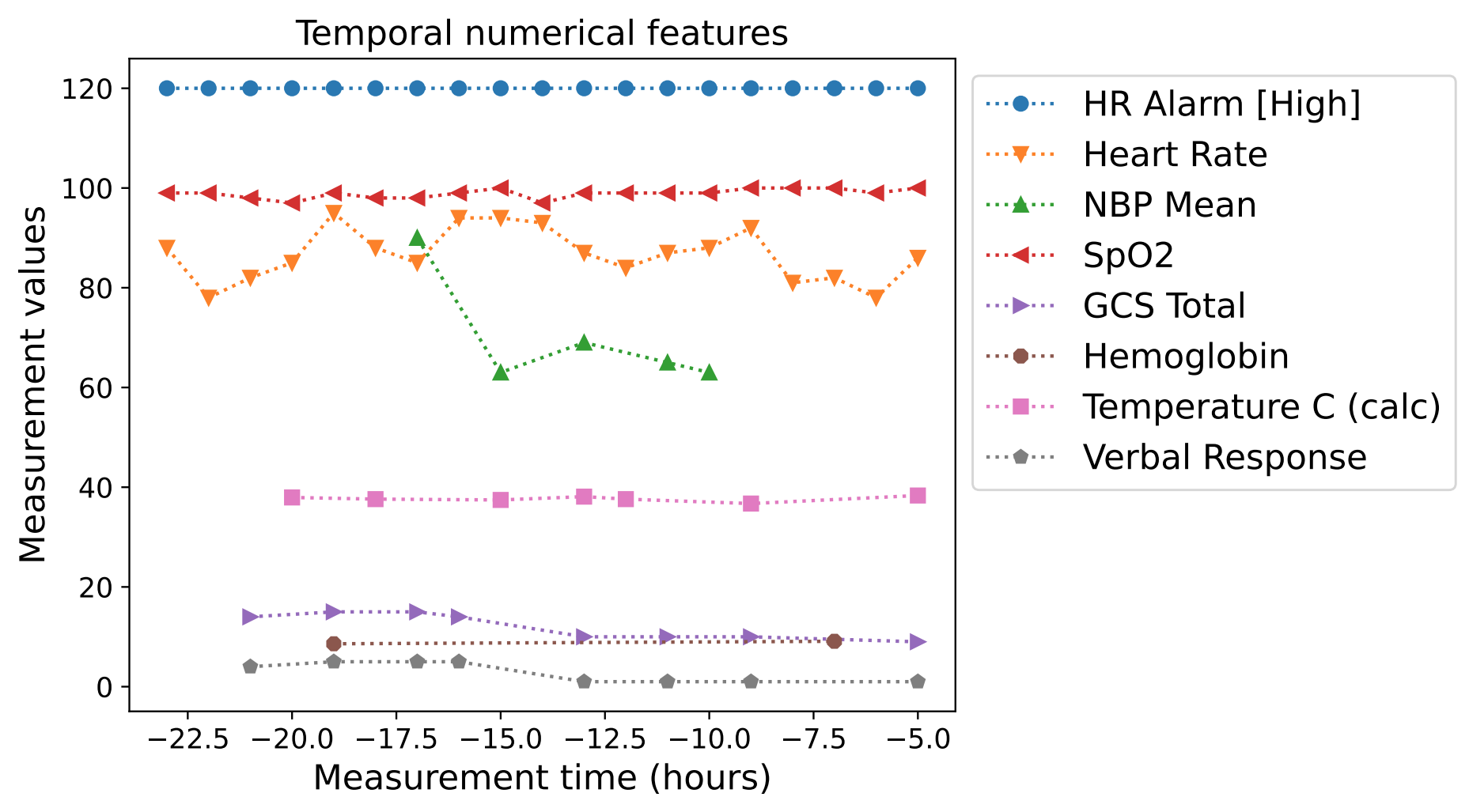 |
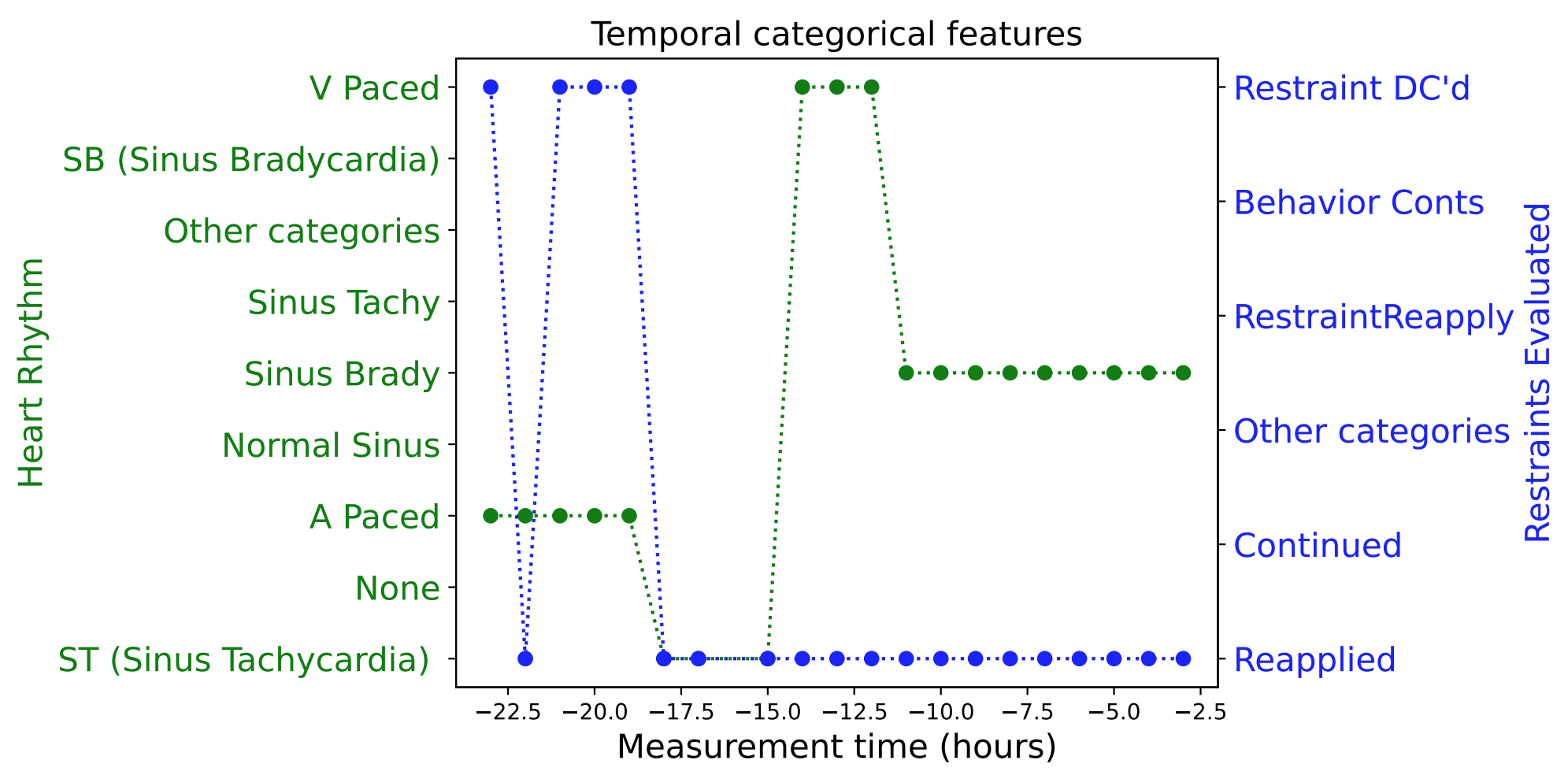 |
| Examples of real EHR data: temporal numerical features (upper) and temporal categorical features (lower). |
EHR-Safe: Synthetic EHR Data Generation Framework
EHR-Safe consists of sequential encoder-decoder architecture and generative adversarial networks (GANs), depicted in the figure below. Because EHR data are heterogeneous (as described above), direct modeling of raw EHR data is challenging for GANs. To circumvent this, we propose utilizing a sequential encoder-decoder architecture, to learn the mapping from the raw EHR data to the latent representations, and vice versa.
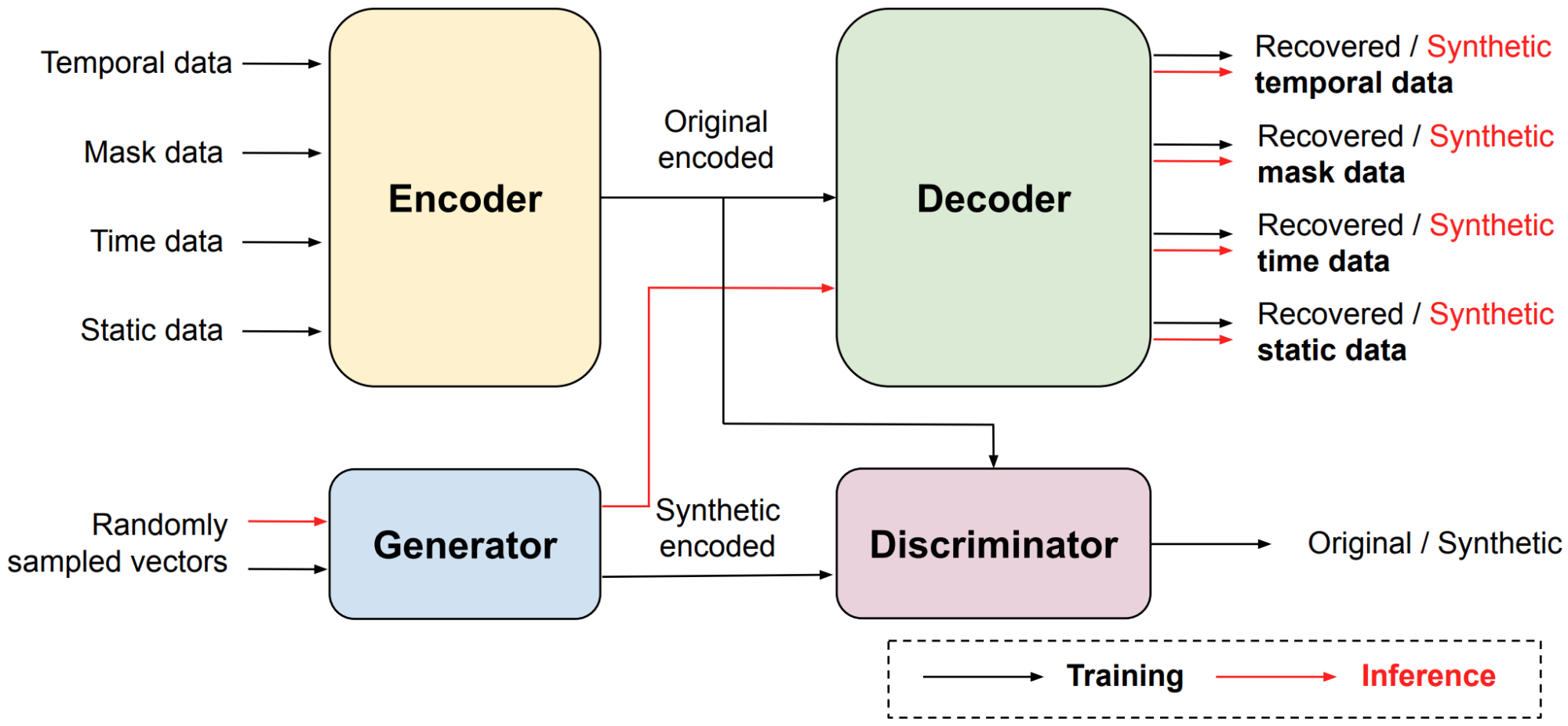 |
| Block diagram of EHR-Safe framework. |
While learning the mapping, esoteric distributions of numerical and categorical features pose a great challenge. For example, some values or numerical ranges might dominate the distribution, but the capability of modeling rare cases is essential. The proposed feature mapping and stochastic normalization (transforming original feature distributions into uniform distributions without information loss) are key to handling such data by converting to distributions for which the training of encoder-decoder and GAN are more stable (details can be found in the paper). The mapped latent representations, generated by the encoder, are then used for GAN training. After training both the encoder-decoder framework and GANs, EHR-Safe can generate synthetic heterogeneous EHR data from any input, for which we feed randomly sampled vectors. Note that only the trained generator and decoders are used for generating synthetic data.
Datasets
We focus on two real-world EHR datasets to showcase the EHR-Safe framework, MIMIC-III and eICU. Both are inpatient datasets that consist of varying lengths of sequences and include multiple numerical and categorical features with missing components.
Fidelity Results
The fidelity metrics focus on the quality of synthetically generated data by measuring the realisticness of the synthetic data. Higher fidelity implies that it is more difficult to differentiate between synthetic and real data. We evaluate the fidelity of synthetic data in terms of multiple quantitative and qualitative analyses.
Visualization
Having similar coverage and avoiding under-representation of certain data regimes are both important for synthetic data generation. As the below t-SNE analyses show, the coverage of the synthetic data (blue) is very similar with the original data (red). With membership inference metrics (will be introduced in the privacy section), we also verify that EHR-Safe does not just memorize the original train data.
 |
| t-SNE analyses on temporal and static data on MIMIC-III (upper) and eICU (lower) datasets. |
Statistical Similarity
We provide quantitative comparisons of statistical similarity between original and synthetic data for each feature. Most statistics are well-aligned between original and synthetic data — for example a measure of the KS statistics, i.e,. the maximum difference in the cumulative distribution function (CDF) between the original and the synthetic data, are mostly lower than 0.03. More detailed tables can be found in the paper. The figure below exemplifies the CDF graphs for original vs. synthetic data for three features — overall they seem very close in most cases.
 |
| CDF graphs of two features between original and synthetic EHR data. Left: Mean Airway Pressure. Right: Minute Volume Alarm. |
Utility
Because one of the most important use cases of synthetic data is enabling ML innovations, we focus on the fidelity metric that measures the ability of models trained on synthetic data to make accurate predictions on real data. We compare such model performance to an equivalent model trained with real data. Similar model performance would indicate that the synthetic data captures the relevant informative content for the task. As one of the important potential use cases of EHR, we focus on the mortality prediction task. We consider four different predictive models: Gradient Boosting Tree Ensemble (GBDT), Random Forest (RF), Logistic Regression (LR), Gated Recurrent Units (GRU).
 |
| Mortality prediction performance with the model trained on real vs. synthetic data. Left: MIMIC-III. Right: eICU. |
In the figure above we see that in most scenarios, training on synthetic vs. real data are highly similar in terms of Area Under Receiver Operating Characteristics Curve (AUC). On MIMIC-III, the best model (GBDT) on synthetic data is only 2.6% worse than the best model on real data; whereas on eICU, the best model (RF) on synthetic data is only 0.9% worse.
Privacy Results
We consider three different privacy attacks to quantify the robustness of the synthetic data with respect to privacy.
- Membership inference attack: An adversary predicts whether a known subject was a present in the training data used for training the synthetic data model.
- Re-identification attack: The adversary explores the probability of some features being re-identified using synthetic data and matching to the training data.
- Attribute inference attack: The adversary predicts the value of sensitive features using synthetic data.
 |
| Privacy risk evaluation across three privacy metrics: membership-inference (top-left), re-identification (top-right), and attribute inference (bottom). The ideal value of privacy risk for membership inference is random guessing (0.5). For re-identification, the ideal case is to replace the synthetic data with disjoint holdout original data. |
The figure above summarizes the results along with the ideal achievable value for each metric. We observe that the privacy metrics are very close to the ideal in all cases. The risk of understanding whether a sample of the original data is a member used for training the model is very close to random guessing; it also verifies that EHR-Safe does not just memorize the original train data. For the attribute inference attack, we focus on the prediction task of inferring specific attributes (e.g., gender, religion, and marital status) from other attributes. We compare prediction accuracy when training a classifier with real data against the same classifier trained with synthetic data. Because the EHR-Safe bars are all lower, the results demonstrate that access to synthetic data does not lead to higher prediction performance on specific features as compared to access to the original data.
Comparison to Alternative Methods
We compare EHR-Safe to alternatives (TimeGAN, RC-GAN, C-RNN-GAN) proposed for time-series synthetic data generation. As shown below, EHR-Safe significantly outperforms each.
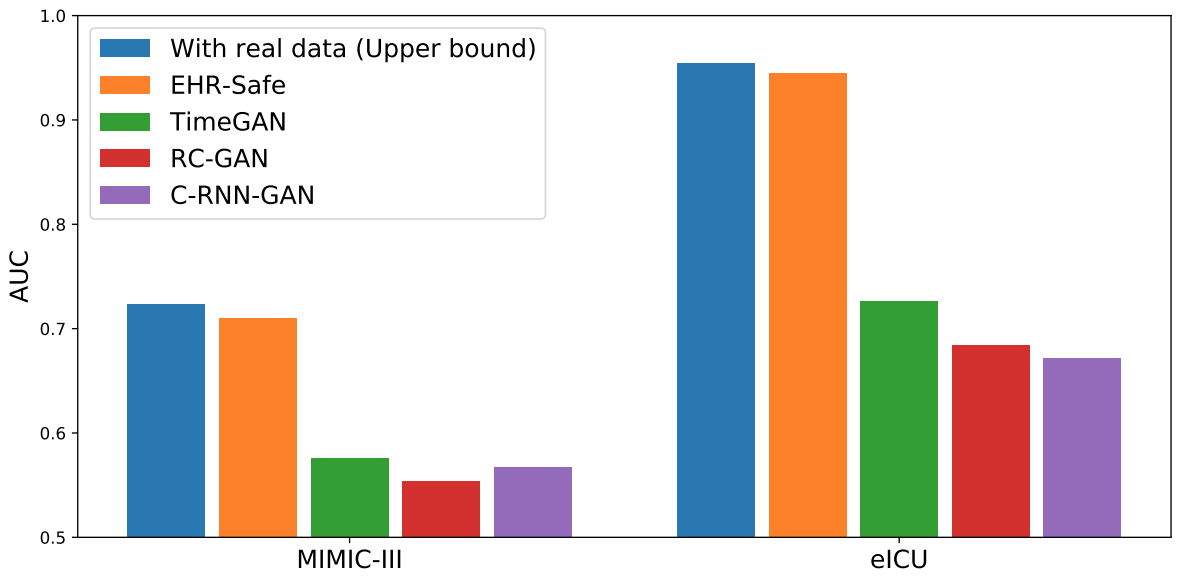 |
| Downstream task performance (AUC) in comparison to alternatives. |
Conclusions
We propose a novel generative modeling framework, EHR-Safe, that can generate highly realistic synthetic EHR data that are robust to privacy attacks. EHR-Safe is based on generative adversarial networks applied to the encoded raw data. We introduce multiple innovations in the architecture and training mechanisms that are motivated by the key challenges of EHR data. These innovations are key to our results that show almost-identical properties with real data (when desired downstream capabilities are considered) with almost-ideal privacy preservation. An important future direction is generative modeling capability for multimodal data, including text and image, as modern EHR data might contain both.
Acknowledgements
We gratefully acknowledge the contributions of Michel Mizrahi, Nahid Farhady Ghalaty, Thomas Jarvinen, Ashwin S. Ravi, Peter Brune, Fanyu Kong, Dave Anderson, George Lee, Arie Meir, Farhana Bandukwala, Elli Kanal, and Tomas Pfister.
 On World Water Day 2023, we’re sharing progress on three partnerships aimed at helping water scarcity and access in India.
On World Water Day 2023, we’re sharing progress on three partnerships aimed at helping water scarcity and access in India.
 On World Water Day 2023, we’re sharing progress on three partnerships aimed at helping water scarcity and access in India.
On World Water Day 2023, we’re sharing progress on three partnerships aimed at helping water scarcity and access in India.


 Google Health helping make people healthier, unveils new features across Search, YouTube and Fitbit as part of its annual Check Up event.
Google Health helping make people healthier, unveils new features across Search, YouTube and Fitbit as part of its annual Check Up event.
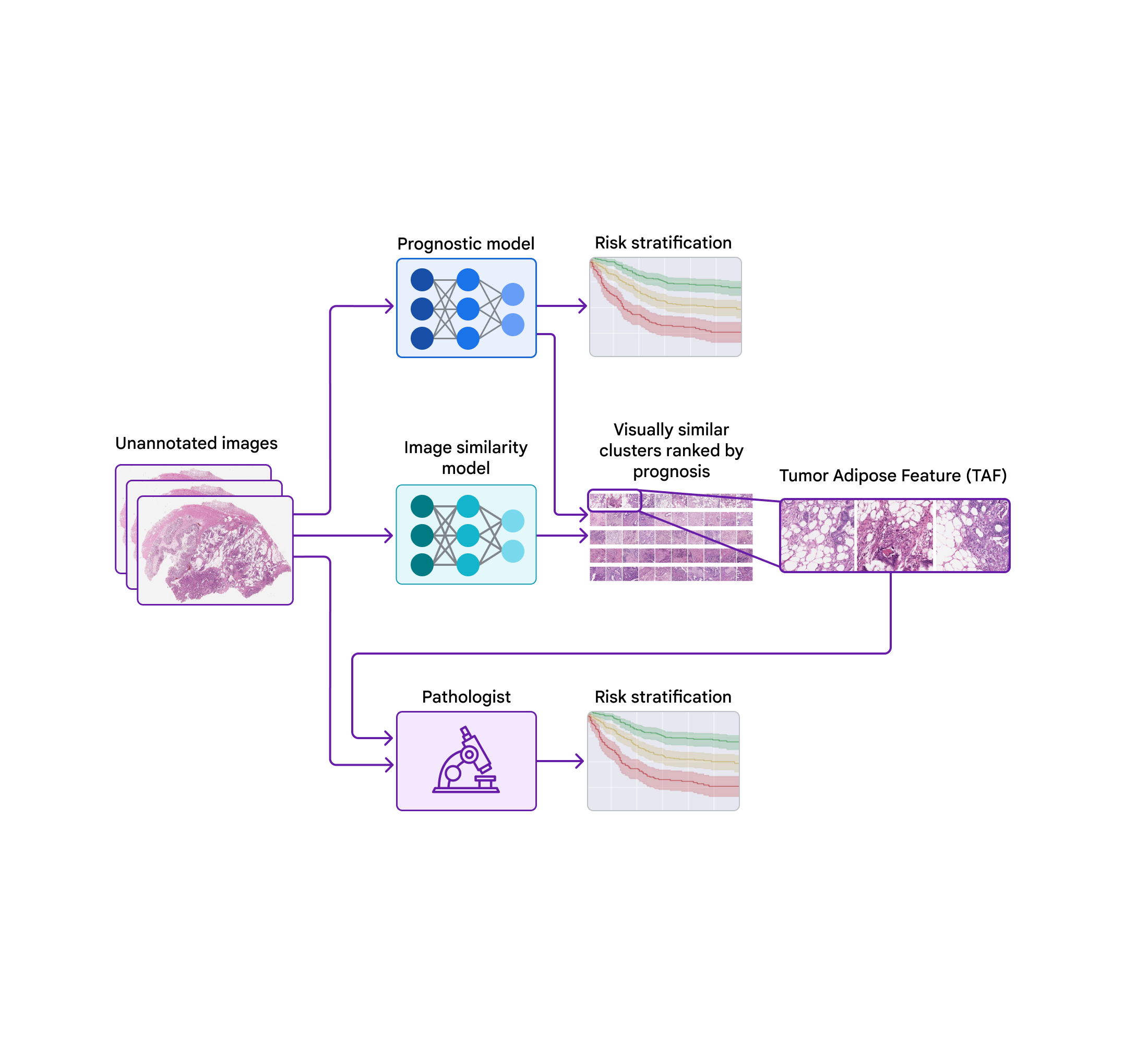
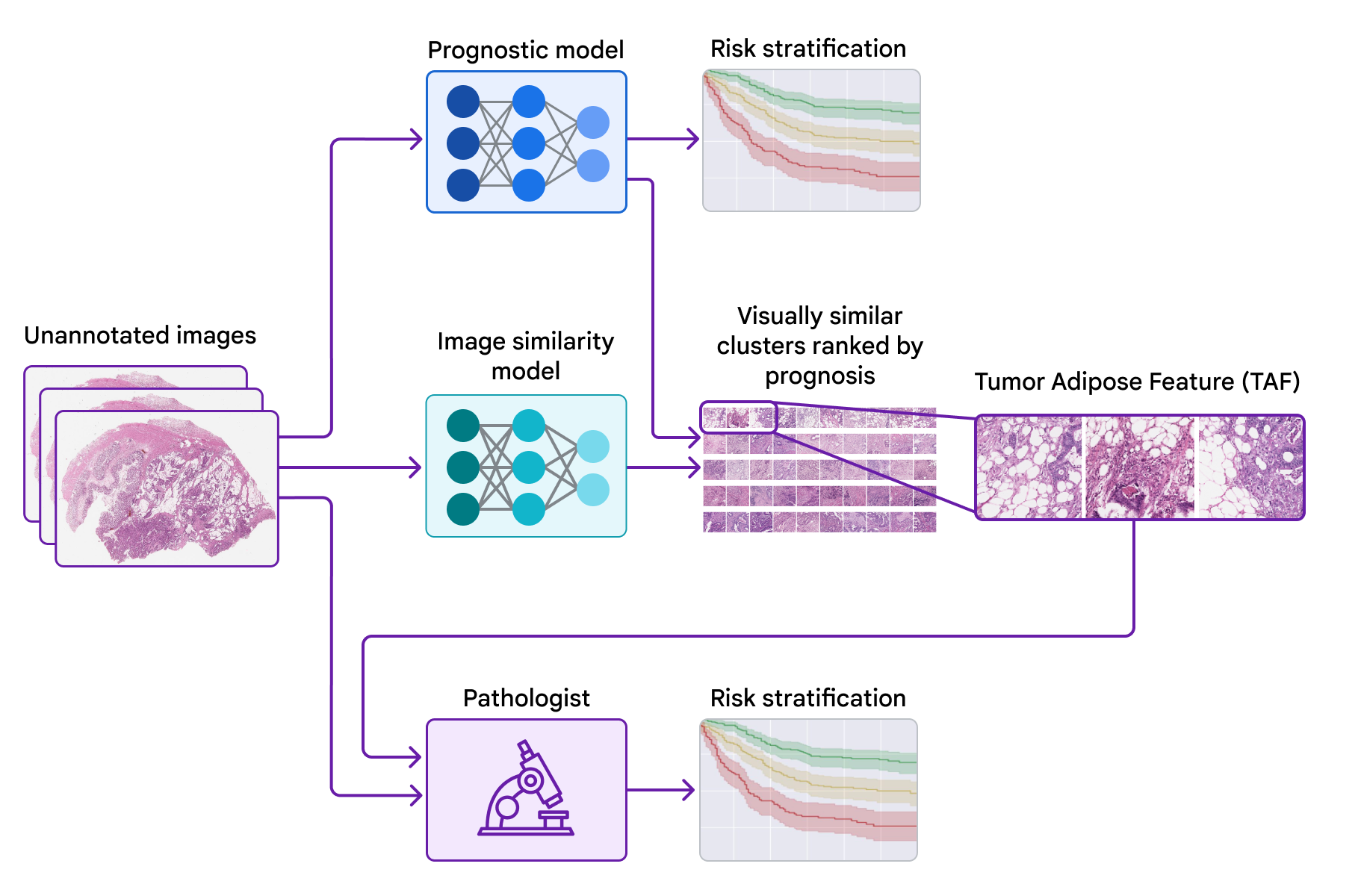
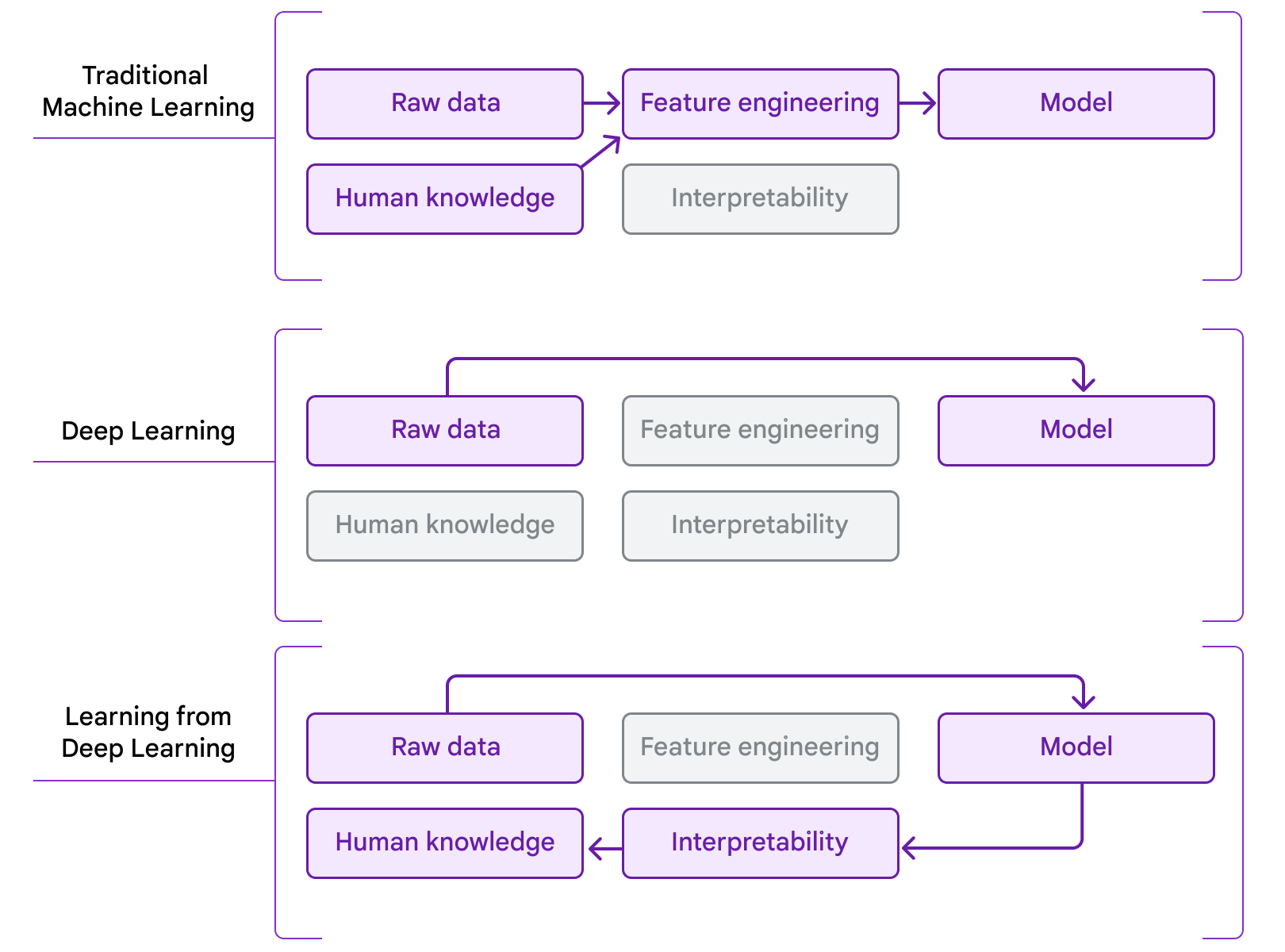
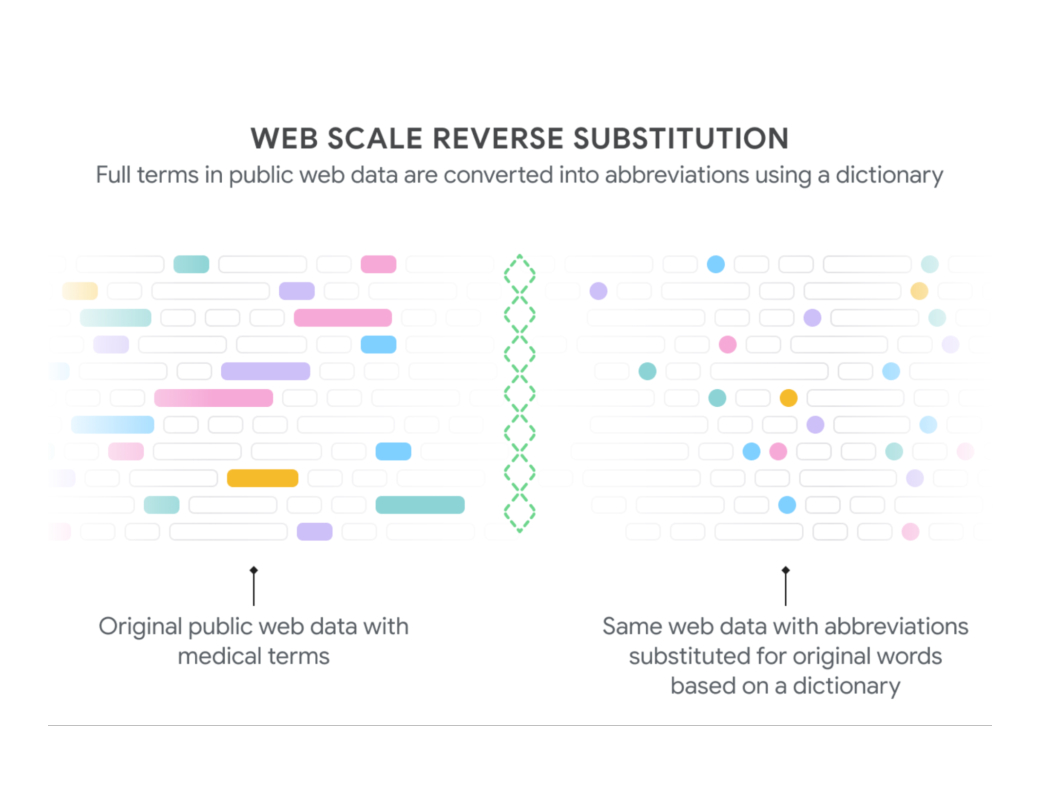
 At The Check Up, we shared updates on our medical LLM research, partnerships and new ways AI can help with disease detection.
At The Check Up, we shared updates on our medical LLM research, partnerships and new ways AI can help with disease detection.
 Today at The Check Up, we introduced Open Health Stack, a suite of open-source building blocks to help developers build digital health apps.
Today at The Check Up, we introduced Open Health Stack, a suite of open-source building blocks to help developers build digital health apps.
 Google Pixel Watch fall detection uses motion sensors and AI to get you the help you need if you take a hard fall.
Google Pixel Watch fall detection uses motion sensors and AI to get you the help you need if you take a hard fall.
 Researchers will receive Cloud credits, funding, Fitbit devices and Fitabase to support research to addresses critical health disparities
Researchers will receive Cloud credits, funding, Fitbit devices and Fitabase to support research to addresses critical health disparities

 How Fitbit, Google Nest and Pixel Watch can help you get better sleep, manage stress and work out smarter in 2023.
How Fitbit, Google Nest and Pixel Watch can help you get better sleep, manage stress and work out smarter in 2023.









