In December 2018, we introduced TF-Ranking, an open-source TensorFlow-based library for developing scalable neural learning-to-rank (LTR) models, which are useful in settings where users expect to receive an ordered list of items in response to their query. LTR models — unlike standard classification models that classify one item at a time — receive an entire list of items as an input, and learn an ordering that maximizes the utility of the entire list. While search and recommendation systems are the most common applications of LTR models, since its release, we have seen TF-Ranking being applied in diverse domains beyond search, including e-commerce, SAT solvers, and smart city planning.
In May 2021, we published a major release of TF-Ranking that enables full support for natively building LTR models using Keras, a high-level API of TensorFlow 2. Our native Keras ranking model has a brand-new workflow design, including a flexible ModelBuilder, a DatasetBuilder to set up training data, and a Pipeline to train the model with the provided dataset. These components make building a customized LTR model easier than ever, and facilitate rapid exploration of new model structures for production and research. If RaggedTensors are your tool of choice, TF-Ranking is now working with them as well. In addition, our most recent release, which incorporates the Orbit training library, contains a long list of advances — the culmination of two and half years of neural LTR research. Below we share a few of the key improvements available in the latest TF-Ranking version.
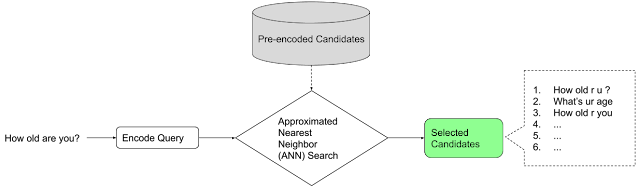
 |
| Workflow to build and train a native Keras ranking model. Blue modules are provided by TF-Ranking, and green modules are customizable. |
Learning-to-Rank with TFR-BERT
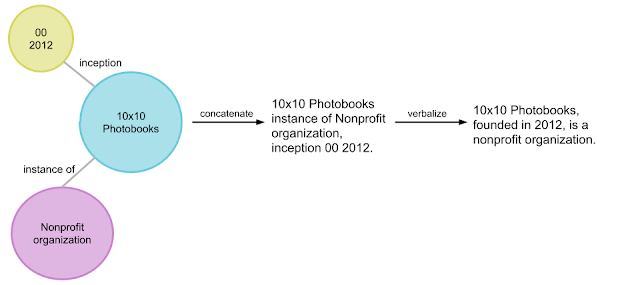
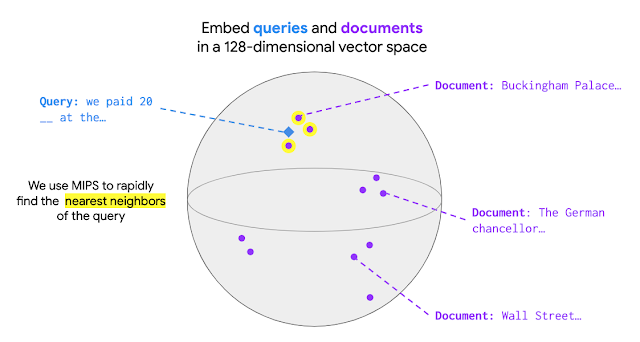
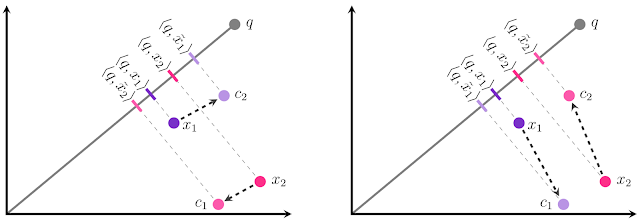
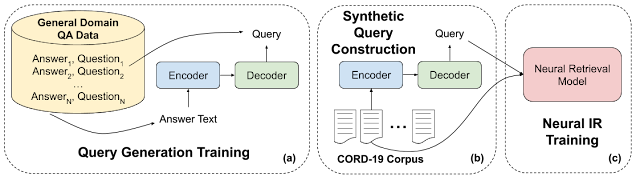
Recently, pretrained language models like BERT have achieved state-of-the-art performance on various language understanding tasks. To capture the expressiveness of these models, TF-Ranking implements a novel TFR-BERT architecture that couples BERT with the power of LTR to optimize the ordering of list inputs. As an example, consider a query and a list of n documents that one might like to rank in response to this query. Instead of learning an independent BERT representation for each <query, document> pair, LTR models apply a ranking loss to jointly learn a BERT representation that maximizes the utility of the entire ranked list with respect to the ground-truth labels.
The figure below illustrates this process. First, we flatten a list of n documents to rank in response to a query into a list <query, document> tuples. These tuples are fed into a pre-trained language model (e.g., BERT). The pooled BERT outputs for the entire document list are then jointly fine-tuned with one of the specialized ranking losses available in TF-Ranking. Our experience shows that this TFR-BERT architecture delivers significant improvements in pretrained language model performance, leading to state-of-the-art performance for several popular ranking tasks, especially when multiple pretrained language models are ensembled. Our users can now get started with TFR-BERT using this simple example.
 |
| An illustration of the TFR-BERT architecture, in which a joint LTR model over a list of n documents is constructed using BERT representations of individual <query, document> pairs. |
Interpretable Learning-to-Rank
Transparency and interpretability are important factors in deploying LTR models in ranking systems that can be involved in determining the outcomes of processes such as loan eligibility assessment, advertisement targeting, or guiding medical treatment decisions. In such cases, the contribution of each individual feature to the final ranking should be examinable and understandable to ensure transparency, accountability and fairness of the outcomes.
One possible way to achieve this is using generalized additive models (GAMs) — intrinsically interpretable machine learning models that are linearly composed of smooth functions of individual features. However, while GAMs have been extensively studied on regression and classification tasks, it is less clear how to apply them in a ranking setting. For instance, while GAMs can be straightforwardly applied to model each individual item in the list, modeling both item interactions and the context in which these items are ranked is a more challenging research problem. To this end, we have developed a neural ranking GAM — an extension of generalized additive models to ranking problems.
Unlike standard GAMs, a neural ranking GAM can take into account both the features of the ranked items and the context features (e.g., query or user profile) to derive an interpretable, compact model. This ensures that not only the contribution of each item-level feature is interpretable, but also the contribution of the context features. For example, in the figure below, using a neural ranking GAM makes visible how distance, price, and relevance, in the context of a given user device, contribute to the final ranking of the hotel. Neural ranking GAMs are now available as a part of TF-Ranking,
Neural Ranking or Gradient Boosting?
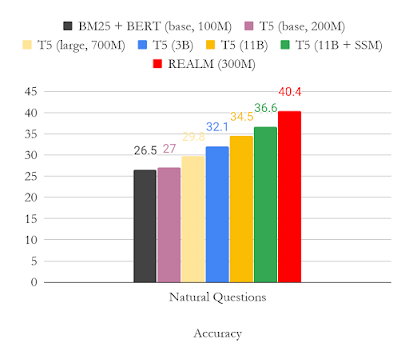
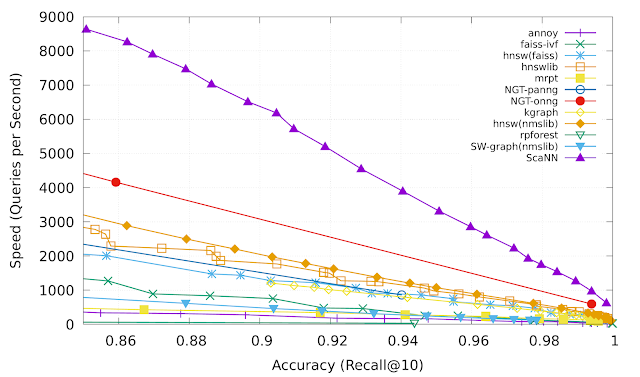
While neural models have achieved state of the art performance in multiple domains, specialized gradient boosted decision trees (GBDTs) like LambdaMART remained the baseline to beat in a variety of open LTR datasets. The success of GBDTs in open datasets is due to several reasons. First, due to their relatively small size, neural models are prone to overfitting on these datasets. Second, since GBDTs partition their input feature space using decision trees, they are naturally more resilient to variations in numerical scales in ranking data, which often contain features with Zipfian or otherwise skewed distributions. However, GBDTs do have their limitations in more realistic ranking scenarios, which often combine both textual and numerical features. For instance, GBDTs cannot be directly applied to large discrete feature spaces, such as raw document text. They are also, in general, less scalable than neural ranking models.
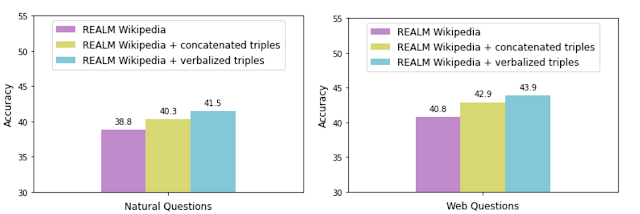
Therefore, since the TF-Ranking release, our team has significantly deepened the understanding of how best to leverage neural models in ranking with numerical features. This culminated in a Data Augmented Self-Attentive Latent Cross (DASALC) model, described in an ICLR 2021 paper, which is the first to establish parity, and in some cases statistically significant improvements, of neural ranking models over strong LambdaMART baselines on open LTR datasets. This achievement is made possible through a combination of techniques, which include data augmentation, neural feature transformation, self-attention for modeling document interactions, listwise ranking loss, and model ensembling similar to boosting in GBDTs. The architecture of the DASALC model was entirely implemented using the TF-Ranking library.
Conclusion
All in all, we believe that the new Keras-based TF-Ranking version will make it easier to conduct neural LTR research and deploy production-grade ranking systems. We encourage everyone to try out the latest version and follow this introductory example for a hands-on experience. While we are very excited about this new release, our research and development journey is far from over, so we will continue to advance our understanding of learning-to-rank problems and share these advances with our users.
Acknowledgements
This project was only possible thanks to the current and past members of the TF-Ranking team: Honglei Zhuang, Le Yan, Rama Pasumarthi, Rolf Jagerman, Zhen Qin, Shuguang Han, Sebastian Bruch, Nathan Cordeiro, Marc Najork and Patrick McGregor. We also extend special thanks to our collaborators from the Tensorflow team: Zhenyu Tan, Goldie Gadde, Rick Chao, Yuefeng Zhou, Hongkun Yu, and Jing Li.