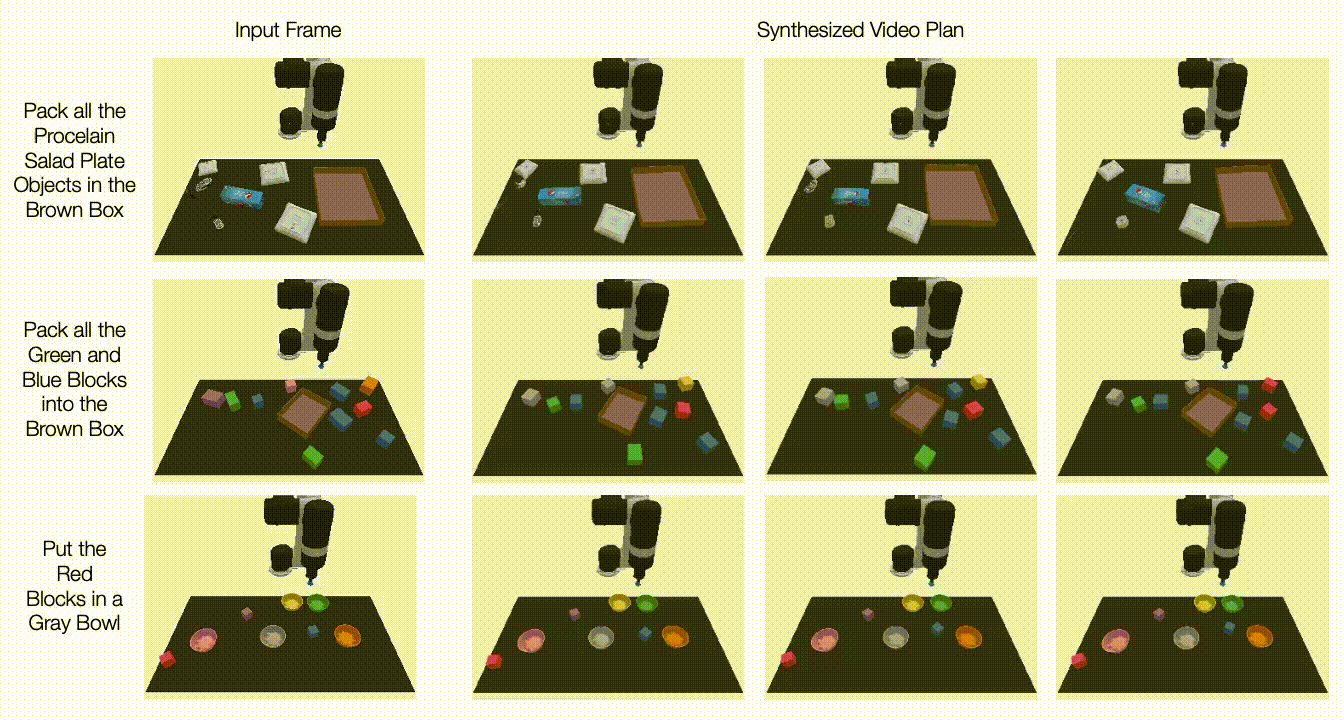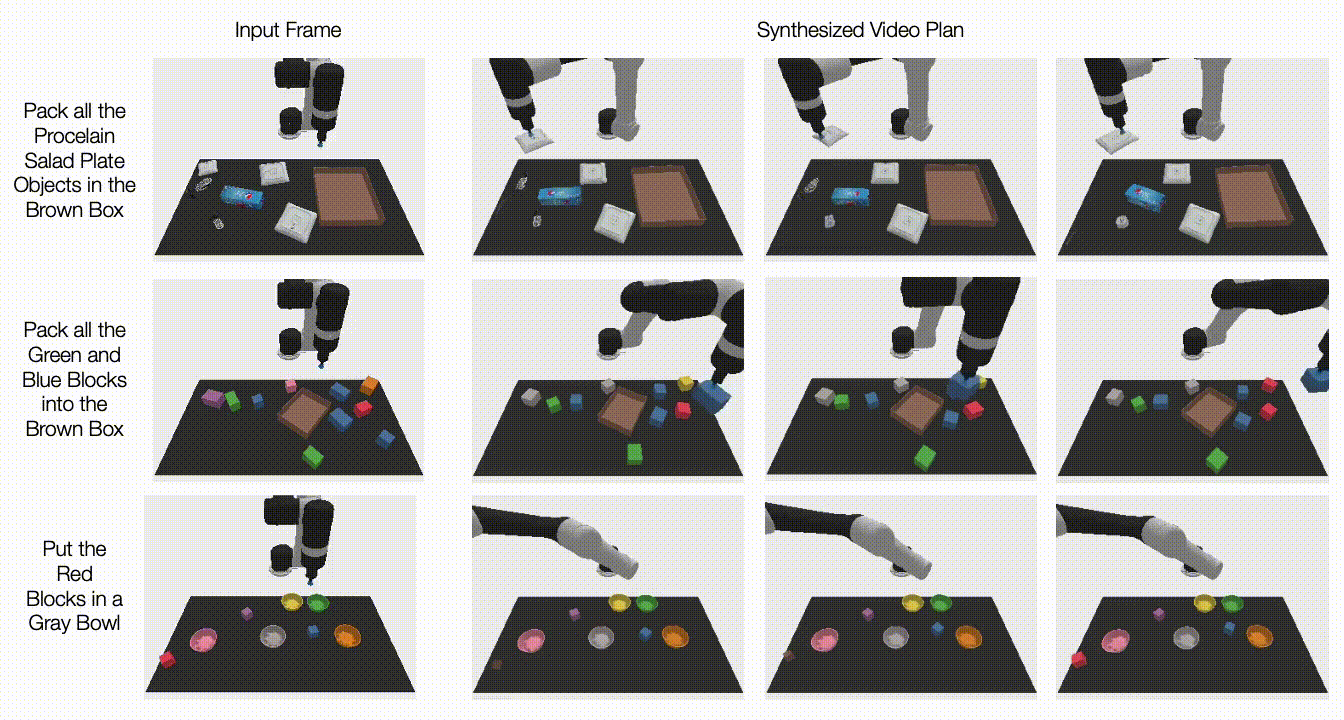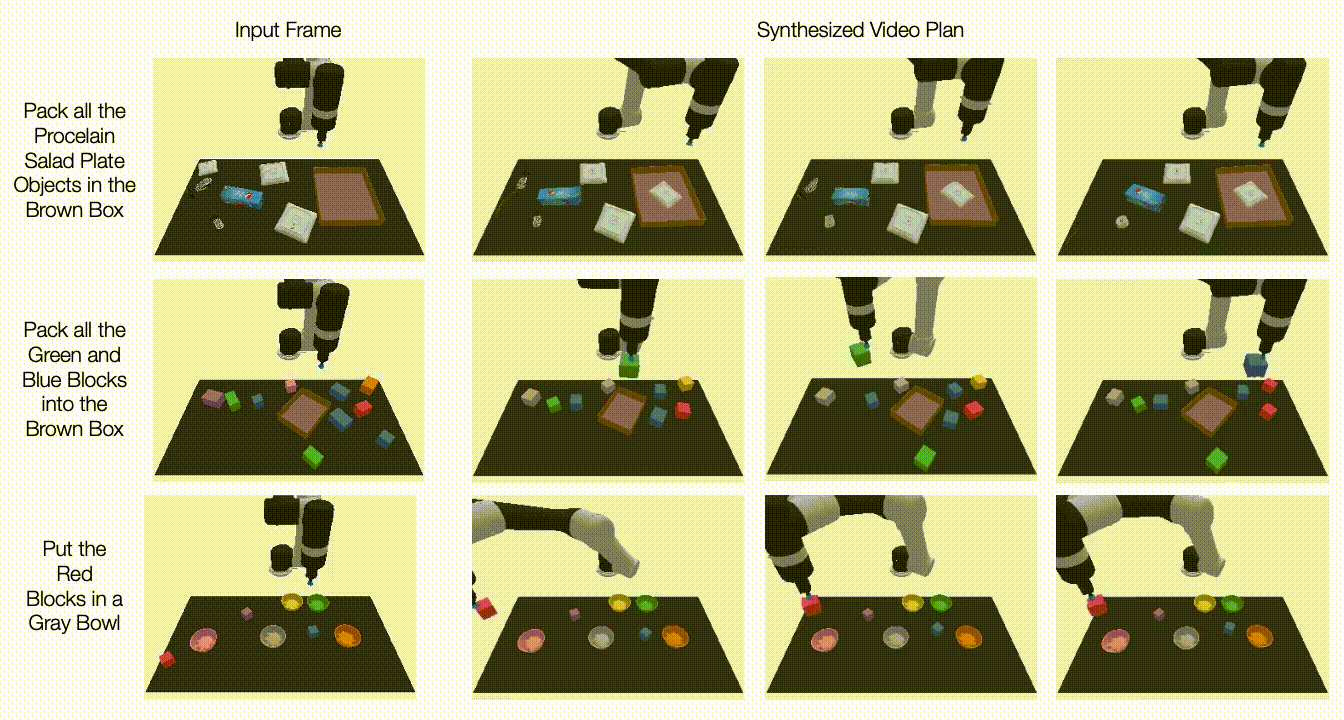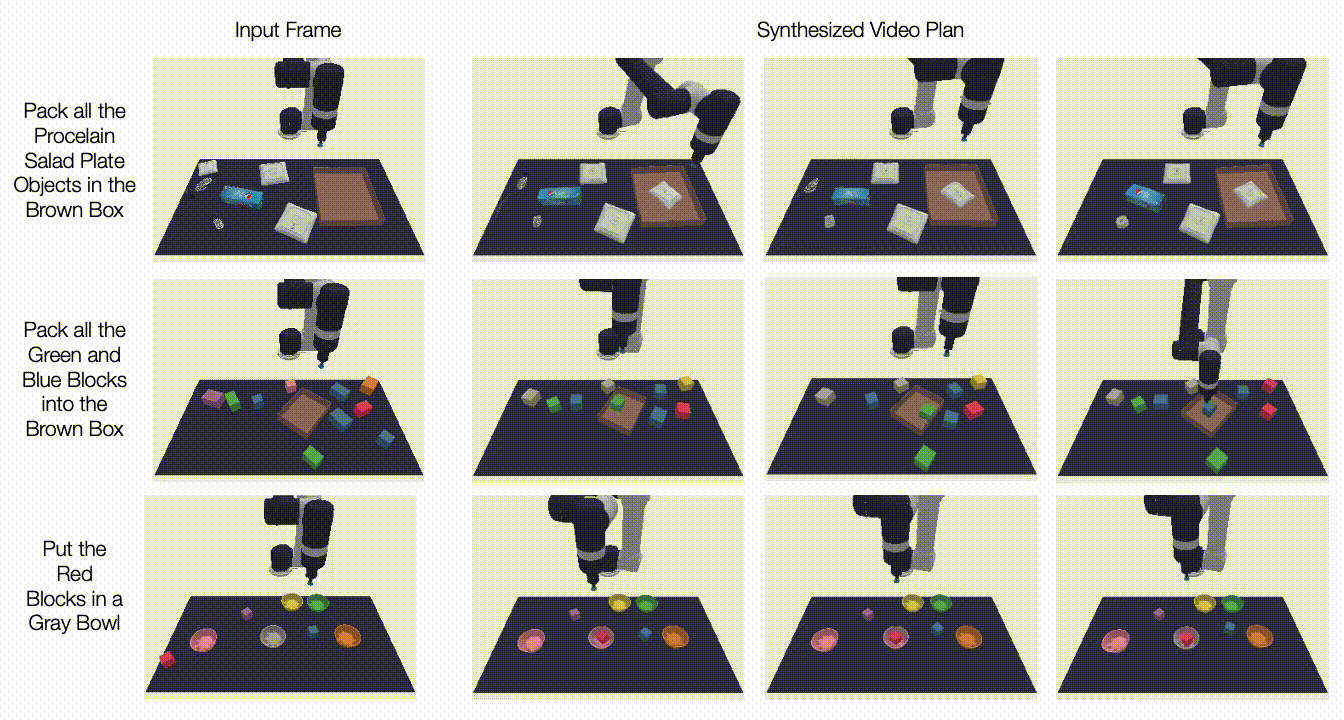
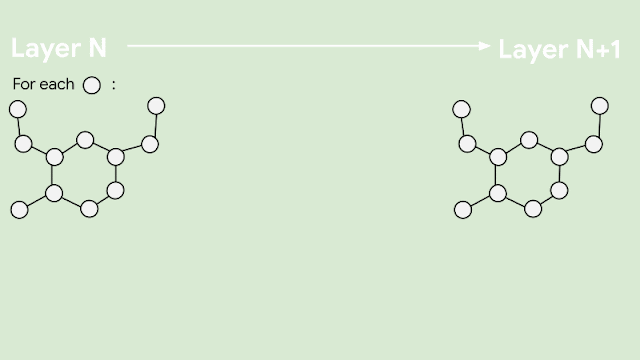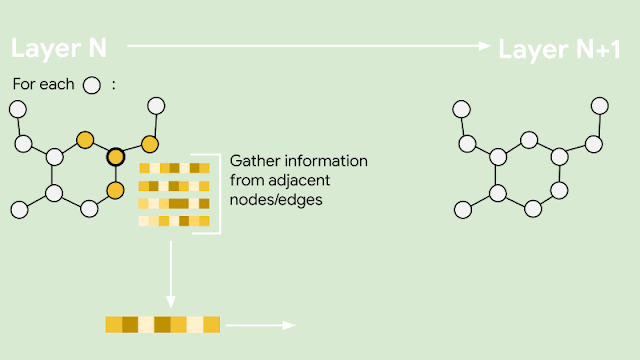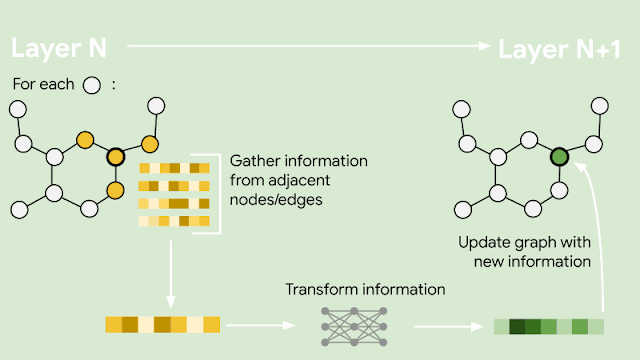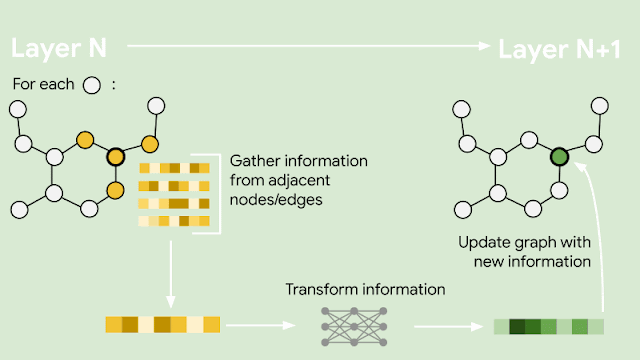
The constantly changing nature of the world around us poses a significant challenge for the development of AI models. Often, models are trained on longitudinal data with the hope that the training data used will accurately represent inputs the model may receive in the future. More generally, the default assumption that all training data are equally relevant often breaks in practice. For example, the figure below shows images from the CLEAR nonstationary learning benchmark, and it illustrates how visual features of objects evolve significantly over a 10 year span (a phenomenon we refer to as slow concept drift), posing a challenge for object categorization models.
 |
| Sample images from the CLEAR benchmark. (Adapted from Lin et al.) |
Alternative approaches, such as online and continual learning, repeatedly update a model with small amounts of recent data in order to keep it current. This implicitly prioritizes recent data, as the learnings from past data are gradually erased by subsequent updates. However in the real world, different kinds of information lose relevance at different rates, so there are two key issues: 1) By design they focus exclusively on the most recent data and lose any signal from older data that is erased. 2) Contributions from data instances decay uniformly over time irrespective of the contents of the data.
In our recent work, “Instance-Conditional Timescales of Decay for Non-Stationary Learning”, we propose to assign each instance an importance score during training in order to maximize model performance on future data. To accomplish this, we employ an auxiliary model that produces these scores using the training instance as well as its age. This model is jointly learned with the primary model. We address both the above challenges and achieve significant gains over other robust learning methods on a range of benchmark datasets for nonstationary learning. For instance, on a recent large-scale benchmark for nonstationary learning (~39M photos over a 10 year period), we show up to 15% relative accuracy gains through learned reweighting of training data.
The challenge of concept drift for supervised learning
To gain quantitative insight into slow concept drift, we built classifiers on a recent photo categorization task, comprising roughly 39M photographs sourced from social media websites over a 10 year period. We compared offline training, which iterated over all the training data multiple times in random order, and continual training, which iterated multiple times over each month of data in sequential (temporal) order. We measured model accuracy both during the training period and during a subsequent period where both models were frozen, i.e., not updated further on new data (shown below). At the end of the training period (left panel, x-axis = 0), both approaches have seen the same amount of data, but show a large performance gap. This is due to catastrophic forgetting, a problem in continual learning where a model’s knowledge of data from early on in the training sequence is diminished in an uncontrolled manner. On the other hand, forgetting has its advantages — over the test period (shown on the right), the continual trained model degrades much less rapidly than the offline model because it is less dependent on older data. The decay of both models’ accuracy in the test period is confirmation that the data is indeed evolving over time, and both models become increasingly less relevant.
 |
| Comparing offline and continually trained models on the photo classification task. |
Time-sensitive reweighting of training data
We design a method combining the benefits of offline learning (the flexibility of effectively reusing all available data) and continual learning (the ability to downplay older data) to address slow concept drift. We build upon offline learning, then add careful control over the influence of past data and an optimization objective, both designed to reduce model decay in the future.
Suppose we wish to train a model, M, given some training data collected over time. We propose to also train a helper model that assigns a weight to each point based on its contents and age. This weight scales the contribution from that data point in the training objective for M. The objective of the weights is to improve the performance of M on future data.
In our work, we describe how the helper model can be meta-learned, i.e., learned alongside M in a manner that helps the learning of the model M itself. A key design choice of the helper model is that we separated out instance- and age-related contributions in a factored manner. Specifically, we set the weight by combining contributions from multiple different fixed timescales of decay, and learn an approximate “assignment” of a given instance to its most suited timescales. We find in our experiments that this form of the helper model outperforms many other alternatives we considered, ranging from unconstrained joint functions to a single timescale of decay (exponential or linear), due to its combination of simplicity and expressivity. Full details may be found in the paper.
Instance weight scoring
The top figure below shows that our learned helper model indeed up-weights more modern-looking objects in the CLEAR object recognition challenge; older-looking objects are correspondingly down-weighted. On closer examination (bottom figure below, gradient-based feature importance assessment), we see that the helper model focuses on the primary object within the image, as opposed to, e.g., background features that may spuriously be correlated with instance age.
 |
| Sample images from the CLEAR benchmark (camera & computer categories) assigned the highest and lowest weights respectively by our helper model. |
 |
| Feature importance analysis of our helper model on sample images from the CLEAR benchmark. |
Results
Gains on large-scale data
We first study the large-scale photo categorization task (PCAT) on the YFCC100M dataset discussed earlier, using the first five years of data for training and the next five years as test data. Our method (shown in red below) improves substantially over the no-reweighting baseline (black) as well as many other robust learning techniques. Interestingly, our method deliberately trades off accuracy on the distant past (training data unlikely to reoccur in the future) in exchange for marked improvements in the test period. Also, as desired, our method degrades less than other baselines in the test period.
 |
| Comparison of our method and relevant baselines on the PCAT dataset. |
Broad applicability
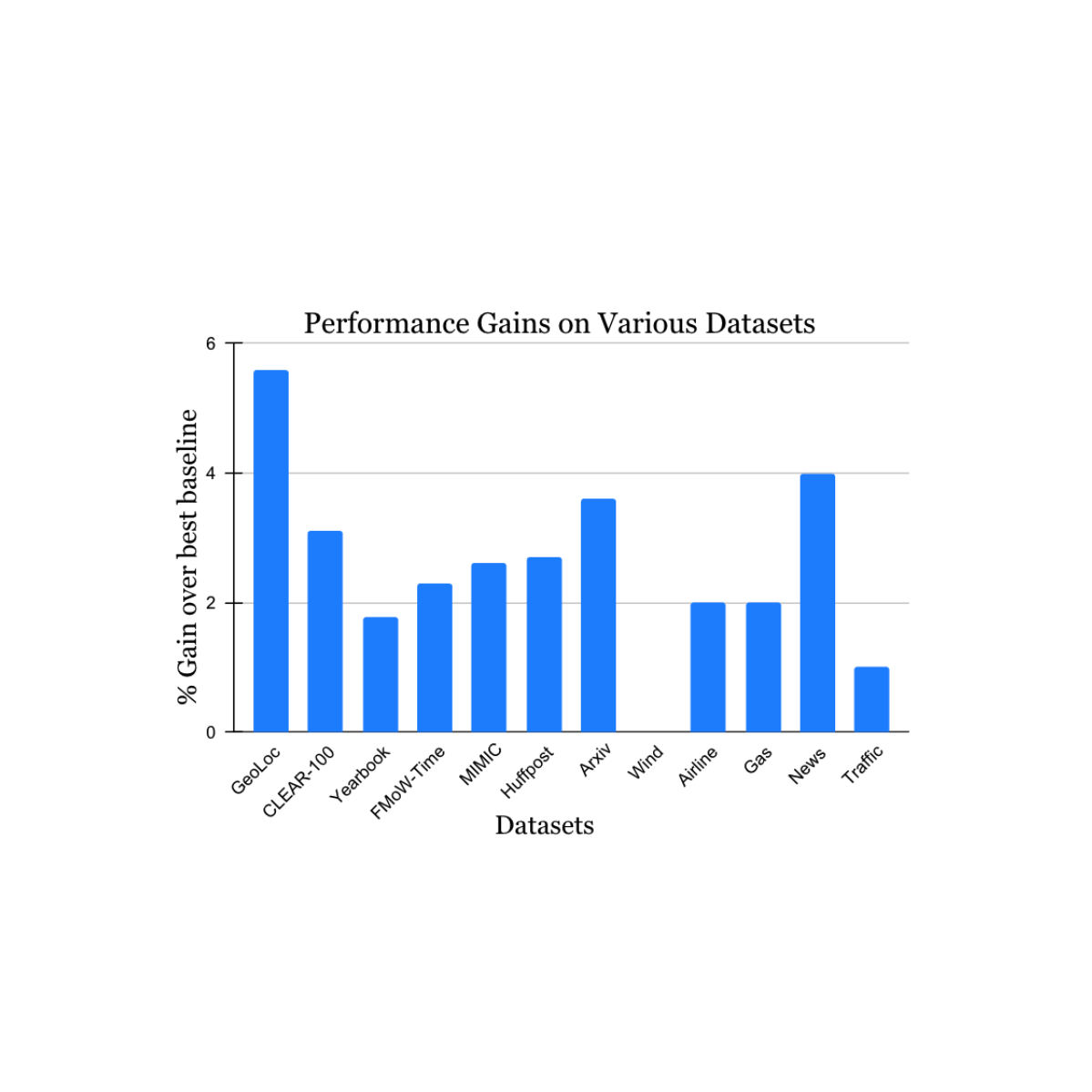
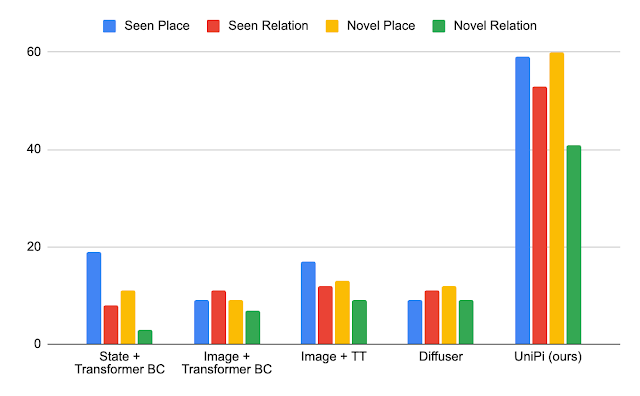
We validated our findings on a wide range of nonstationary learning challenge datasets sourced from the academic literature (see 1, 2, 3, 4 for details) that spans data sources and modalities (photos, satellite images, social media text, medical records, sensor readings, tabular data) and sizes (ranging from 10k to 39M instances). We report significant gains in the test period when compared to the nearest published benchmark method for each dataset (shown below). Note that the previous best-known method may be different for each dataset. These results showcase the broad applicability of our approach.
 |
| Performance gain of our method on a variety of tasks studying natural concept drift. Our reported gains are over the previous best-known method for each dataset. |
Extensions to continual learning
Finally, we consider an interesting extension of our work. The work above described how offline learning can be extended to handle concept drift using ideas inspired by continual learning. However, sometimes offline learning is infeasible — for example, if the amount of training data available is too large to maintain or process. We adapted our approach to continual learning in a straightforward manner by applying temporal reweighting within the context of each bucket of data being used to sequentially update the model. This proposal still retains some limitations of continual learning, e.g., model updates are performed only on most-recent data, and all optimization decisions (including our reweighting) are only made over that data. Nevertheless, our approach consistently beats regular continual learning as well as a wide range of other continual learning algorithms on the photo categorization benchmark (see below). Since our approach is complementary to the ideas in many baselines compared here, we anticipate even larger gains when combined with them.
 |
| Results of our method adapted to continual learning, compared to the latest baselines. |
Conclusion
We addressed the challenge of data drift in learning by combining the strengths of previous approaches — offline learning with its effective reuse of data, and continual learning with its emphasis on more recent data. We hope that our work helps improve model robustness to concept drift in practice, and generates increased interest and new ideas in addressing the ubiquitous problem of slow concept drift.
Acknowledgements
We thank Mike Mozer for many interesting discussions in the early phase of this work, as well as very helpful advice and feedback during its development.