With the increasing ability to consistently and accurately process large amounts of data, particularly visual data, computer-aided diagnostic systems are more frequently being used to assist physicians in their work. This, in turn, can lead to meaningful improvements in health care. An example of where this could be especially useful is in the diagnosis and treatment of colorectal cancer (CRC), which is especially deadly and results in over 900K deaths per year, globally. CRC originates in small pre-cancerous lesions in the colon, called polyps, the identification and removal of which is very successful in preventing CRC-related deaths.
The standard procedure used by gastroenterologists (GIs) to detect and remove polyps is the colonoscopy, and about 19 million such procedures are performed annually in the US alone. During a colonoscopy, the gastroenterologist uses a camera-containing probe to check the intestine for pre-cancerous polyps and early signs of cancer, and removes tissue that looks worrisome. However, complicating factors, such as incomplete detection (in which the polyp appears within the field of view, but is missed by the GI, perhaps due to its size or shape) and incomplete exploration (in which the polyp does not appear in the camera’s field of view), can lead to a high fraction of missed polyps. In fact, studies suggest that 22%–28% of polyps are missed during colonoscopies, of which 20%–24% have the potential to become cancerous (adenomas).
Today, we are sharing progress made in using machine learning (ML) to help GIs fight colorectal cancer by making colonoscopies more effective. In “Detection of Elusive Polyps via a Large Scale AI System”, we present an ML model designed to combat the problem of incomplete detection by helping the GI detect polyps that are within the field of view. This work adds to our previously published work that maximizes the coverage of the colon during the colonoscopy by flagging for GI follow-up areas that may have been missed. Using clinical studies, we show that these systems significantly improve polyp detection rates.
Incomplete Exploration
To help the GI detect polyps that are outside the field of view, we previously developed an ML system that reduces the rate of incomplete exploration by estimating the fractions of covered and non-covered regions of a colon during a colonoscopy. This earlier work uses computer vision and geometry in a technique we call colonoscopy coverage deficiency via depth, to compute segment-by-segment coverage for the colon. It does so in two phases: first computing depth maps for each frame of the colonoscopy video, and then using these depth maps to compute the coverage in real time.
 |
| The ML system computes a depth image (middle) from a single RGB image (left). Then, based on the computation of depth images for a video sequence, it calculates local coverage (right), and detects where the coverage has been deficient and a second look is required (blue color indicates observed segments where red indicates uncovered ones). You can learn more about this work in our previous blog post. |
This segment-by-segment work yields the ability to estimate what fraction of the current segment has been covered. The helpfulness of such functionality is clear: during the procedure itself, a physician may be alerted to segments with deficient coverage, and can immediately return to review these areas, potentially reducing the rates of missed polyps due to incomplete exploration.
Incomplete Detection
In our most recent paper, we look into the problem of incomplete detection. We describe an ML model that aids a GI in detecting polyps that are within the field of view, so as to reduce the rate of incomplete detection. We developed a system that is based on convolutional neural networks (CNN) with an architecture that combines temporal logic with a single frame detector, resulting in more accurate detection.
This new system has two principal advantages. The first is that the system improves detection performance by reducing the number of false negatives detections of elusive polyps, those polyps that are particularly difficult for GIs to detect. The second advantage is the very low false positive rate of the system. This low false positive rate makes these systems more likely to be adopted in the clinic.
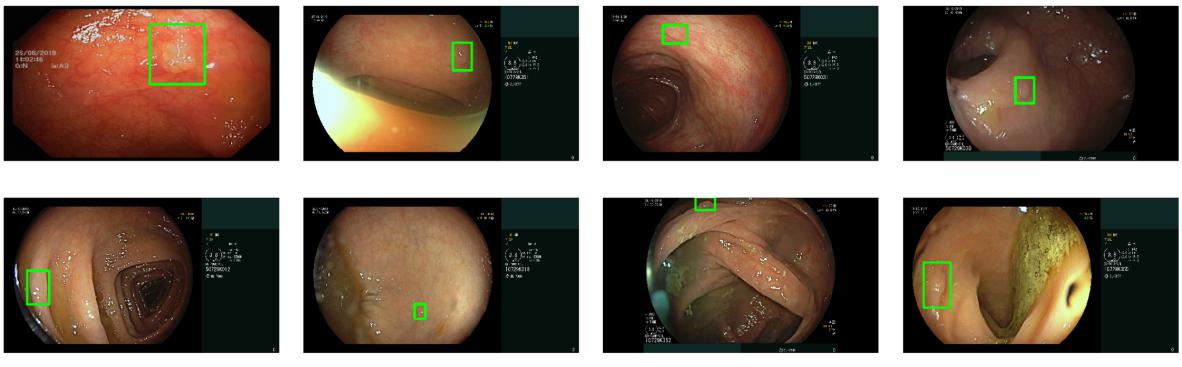 |
| Examples of the variety of polyps detected by the ML system. |
We trained the system on 3600 procedures (86M video frames) and tested it on 1400 procedures (33M frames). All the videos and metadata were de-identified. The system detected 97% of the polyps (i.e., it yielded 97% sensitivity) at 4.6 false alarms per procedure, which is a substantial improvement over previously published results. Of the false alarms, follow-up review showed that some were, in fact, valid polyp detections, indicating that the system was able to detect polyps that were missed by the performing endoscopist and by those who annotated the data. The performance of the system on these elusive polyps suggests its generalizability in that the system has learned to detect examples that were initially missed by all who viewed the procedure.
We evaluated the system performance on polyps that are in the field of view for less than five seconds, which makes them more difficult for the GI to detect, and for which models typically have much lower sensitivity. In this case the system attained a sensitivity that is about three times that of the sensitivity that the original procedure achieved. When the polyps were present in the field of view for less than 2 seconds, the difference was even more stark — the system exhibited a 4x improvement in sensitivity.
It is also interesting to note that the system is fairly insensitive to the choice of neural network architecture. We used two architectures: RetinaNet and LSTM-SSD. RetinaNet is a leading technique for object detection on static images (used for video by applying it to frames in a consecutive fashion). It is one of the top performers on a variety of benchmarks, given a fixed computational budget, and is known for balancing speed of computation with accuracy. LSTM-SSD is a true video object detection architecture, which can explicitly account for the temporal character of the video (e.g., temporal consistency of detections, ability to deal with blur and fast motion, etc.). It is known for being robust and very computationally lightweight and can therefore run on less expensive processors. Comparable results were also obtained on the much heavier Faster R-CNN architecture. The fact that results are similar across different architectures implies that one can choose the network meeting the available hardware specifications.
Prospective Clinical Research Study
As part of the research reported in our detection paper we ran a clinical validation on 100 procedures in collaboration with Shaare Zedek Medical Center in Jerusalem, where our system was used in real time to help GIs. The system helped detect an average of one polyp per procedure that would have otherwise been missed by the GI performing the procedure, while not missing any of the polyps detected by the GIs, and with 3.8 false alarms per procedure. The feedback from the GIs was consistently positive.
We are encouraged by the potential helpfulness of this system for improving polyp detection, and we look forward to working together with the doctors in the procedure room to further validate this research.
Acknowledgements
The research was conducted by teams from Google Health and Google Research, Israel with support from Verily Life Sciences, and in collaboration with Shaare Zedek Medical Center. Verily is advancing this research via a newly established center in Israel, led by Ehud Rivlin. This research was conducted by Danny Veikherman, Tomer Golany, Dan M. Livovsky, Amit Aides, Valentin Dashinsky, Nadav Rabani, David Ben Shimol, Yochai Blau, Liran Katzir, Ilan Shimshoni, Yun Liu, Ori Segol, Eran Goldin, Greg Corrado, Jesse Lachter, Yossi Matias, Ehud Rivlin, and Daniel Freedman. Our appreciation also goes to several institutions and GIs who provided advice along the way and tested our system prototype. We would like to thank all of our team members and collaborators who worked on this project with us, including: Chen Barshai, Nia Stoykova, and many others.